Information
- Publication Type: Journal Paper with Conference Talk
- Workgroup(s)/Project(s):
- Date: June 2019
- Journal: Computer Graphics Forum
- Volume: 38
- Number: 3
- Location: Porto, Portugal
- Lecturer: Jan Byska
- Event: EuroVis 2019
- DOI: 10.1111/cgf.13701
- Call for Papers: Call for Paper
- Conference date: 3. June 2019 – 7. June 2019
- Pages: 441 – 453
- Keywords: scientific visualization, user centered design
Abstract
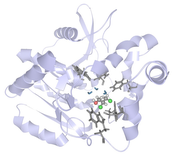
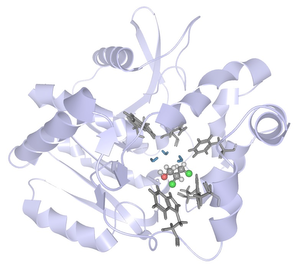
Analyzing molecular dynamics (MD) simulations is a key aspect to understand protein dynamics and function. With increasing computational power, it is now possible to generate very long and complex simulations, which are cumbersome to explore using traditional 3D animations of protein movements. Guided by requirements derived from multiple focus groups with protein engineering experts, we designed and developed a novel interactive visual analysis approach for long and crowded MD simulations. In this approach, we link a dynamic 3D focus+context visualization with a 2D chart of time series data to guide the detection and navigation towards important spatio-temporal events. The 3D visualization renders elements of interest in more detail and increases the temporal resolution dependent on the time series data or the spatial region of interest. In case studies with different MD simulation data sets and research questions, we found that the proposed visual analysis approach facilitates exploratory analysis to generate, confirm, or reject hypotheses about causalities. Finally, we derived design guidelines for interactive visual analysis of complex MD simulation data.Additional Files and Images
Weblinks
BibTeX
@article{byska-2019-mdfc,
title = "Analysis of Long Molecular Dynamics Simulations Using
Interactive Focus+Context Visualization",
author = "Jan Byska and Thomas Trautner and Sergio Marques and Jiri
Damborsky and Barbora Kozlikova and Manuela Waldner",
year = "2019",
abstract = "Analyzing molecular dynamics (MD) simulations is a key
aspect to understand protein dynamics and function. With
increasing computational power, it is now possible to
generate very long and complex simulations, which are
cumbersome to explore using traditional 3D animations of
protein movements. Guided by requirements derived from
multiple focus groups with protein engineering experts, we
designed and developed a novel interactive visual analysis
approach for long and crowded MD simulations. In this
approach, we link a dynamic 3D focus+context visualization
with a 2D chart of time series data to guide the detection
and navigation towards important spatio-temporal events. The
3D visualization renders elements of interest in more detail
and increases the temporal resolution dependent on the time
series data or the spatial region of interest. In case
studies with different MD simulation data sets and research
questions, we found that the proposed visual analysis
approach facilitates exploratory analysis to generate,
confirm, or reject hypotheses about causalities. Finally, we
derived design guidelines for interactive visual analysis of
complex MD simulation data.",
month = jun,
journal = "Computer Graphics Forum",
volume = "38",
number = "3",
doi = "10.1111/cgf.13701",
pages = "441--453",
keywords = "scientific visualization, user centered design",
URL = "https://www.cg.tuwien.ac.at/research/publications/2019/byska-2019-mdfc/",
}

 paper
paper video
video