Information
- Publication Type: Bachelor Thesis
- Workgroup(s)/Project(s):
- Date: September 2014
- Matrikelnummer: 1125229
- First Supervisor:
Abstract
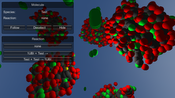
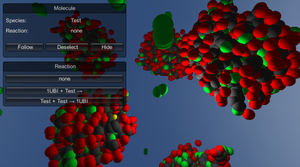
CellUnity is a tool for interactive visualization of molecular reactions using the Unity game engine. Current mesoscale visualizations commonly utilize the results of particle-based simulations, which account for spatial information of each single particle and are supposed to mimic a realistic behavior of the metabolites. However, this approach employs stochastic simulation methods which do not offer any control over the visualized output. CellUnity, on the other hand, exploits the results of deterministic simulations which are purely quantitative and in that way offering full user control over the spatial locations of the reactions. The user is able to trigger reactions on demand instead of having to wait or search for a specific type of reaction event, while the quantities of displayed molecules would still be in accordance with real scientific data. CellUnity exploits the simulation results in real time and allows the user to freely modify simulation parameters while the system is running. The tool was realized in Unity, a cross-platform game engine that also comprises a free version with adequate functionality and therefore enables easy deployment of the project.Additional Files and Images
Weblinks
No further information available.BibTeX
@bachelorsthesis{Gehrer_Daniel_CUI,
title = "CellUnity an Interactive Tool for Illustrative Visualization
of Molecular Reactions",
author = "Daniel Gehrer",
year = "2014",
abstract = "CellUnity is a tool for interactive visualization of
molecular reactions using the Unity game engine. Current
mesoscale visualizations commonly utilize the results of
particle-based simulations, which account for spatial
information of each single particle and are supposed to
mimic a realistic behavior of the metabolites. However, this
approach employs stochastic simulation methods which do not
offer any control over the visualized output. CellUnity, on
the other hand, exploits the results of deterministic
simulations which are purely quantitative and in that way
offering full user control over the spatial locations of the
reactions. The user is able to trigger reactions on demand
instead of having to wait or search for a specific type of
reaction event, while the quantities of displayed molecules
would still be in accordance with real scientific data.
CellUnity exploits the simulation results in real time and
allows the user to freely modify simulation parameters while
the system is running. The tool was realized in Unity, a
cross-platform game engine that also comprises a free
version with adequate functionality and therefore enables
easy deployment of the project.",
month = sep,
address = "Favoritenstrasse 9-11/E193-02, A-1040 Vienna, Austria",
school = "Institute of Computer Graphics and Algorithms, Vienna
University of Technology ",
URL = "https://www.cg.tuwien.ac.at/research/publications/2014/Gehrer_Daniel_CUI/",
}
 Bakk Thesis
Bakk Thesis

